What genetic characteristics characterized the Spanish flu, which killed between 50 and 100 million people from 1918 onwards? Using tissue samples collected in Germany in 1918, researchers have now reconstructed the genomes of influenza viruses of the time and compared them with other historical samples from America and with the seasonal influenza viruses circulating today. Thus, the H1N1 virus responsible for the 1918 pandemic showed distinctive changes that likely made it more virulent than previous strains. In addition, it showed clear regional diversity – and may form the basis for all seasonal H1N1 strains circulating today.
The 1918 flu pandemic, known as the Spanish Flu, killed an estimated 50 to 100 million people worldwide. It peaked in the fall of 1918, and another wave followed in the spring of 1919. Although it was suspected at the beginning of the pandemic that it was caused by a virus, its viral origin was not established until 1930. In the 1990s, research on tissue samples in the Pandemic era Virus as an influenza A virus of the H1N1 subtype. Since then, two complete genomes of the virus have been isolated from American samples, as have many other incomplete samples. However, genetic material from that time is scarce, so many questions remain unanswered.
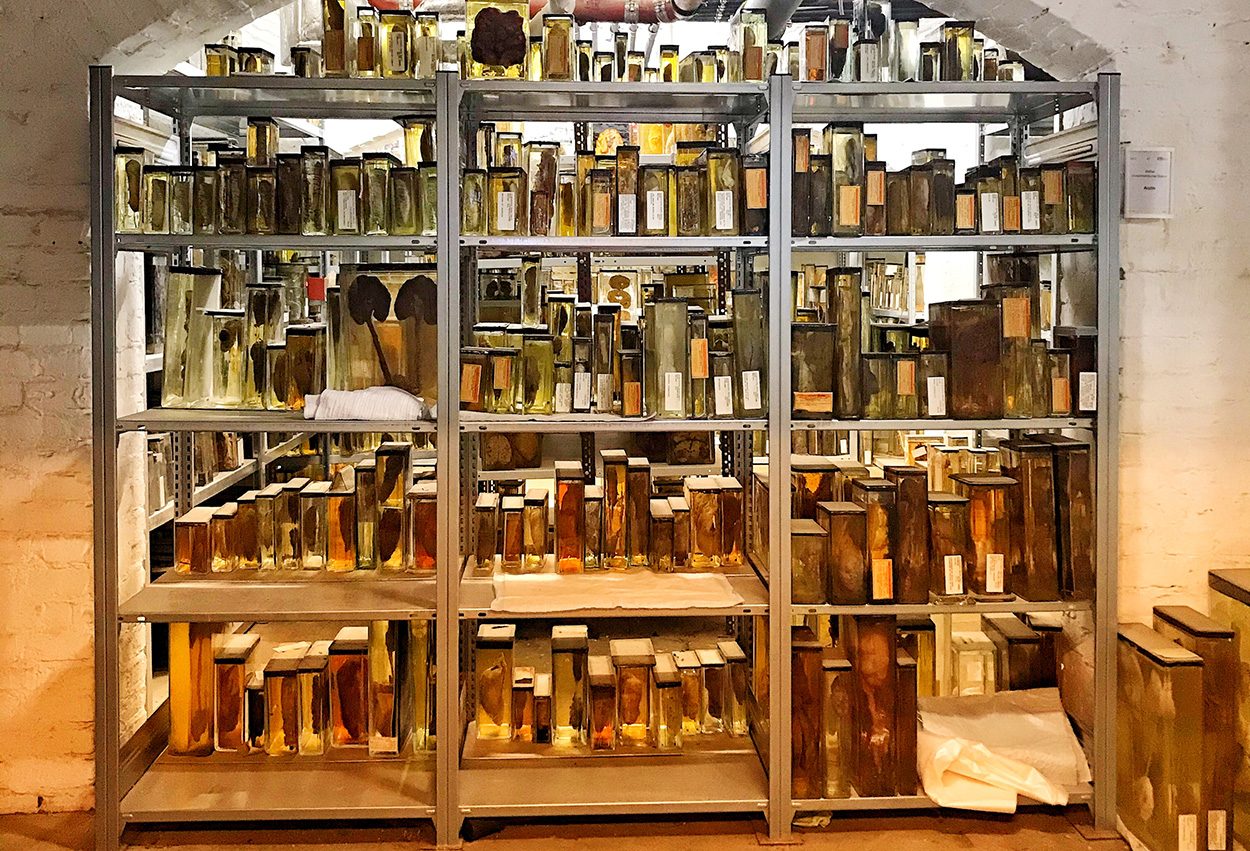
Find clues in medical history museums
In search of other suitable specimens, a team led by Livia Patrono of the Robert Koch Institute (RKI) in Berlin analyzed a total of 13 lung specimens from 1900 and 1931 kept at the Charity Museum in Berlin and the Natural History Museum in Vienna. Six of the lungs preserved in formalin were from the time of the pandemic. Patrono and her team analyzed the genomes of all the samples and actually found influenza A virus RNA in three of them. “All samples from 1918 and the lungs show that bronchopneumonia was present,” the researchers said.
Two samples of influenza A viruses established in Berlin were collected in June 1918, that is, in the first phase of the epidemic. The researchers were able to sequence sections of the viral genome from these samples. In the third sample, collected in Munich in 1918, influenza RNA was so well preserved that researchers were able to sequence the entire genome. Patrono and her team compared the genomic data to already known genome sequences from the United States, one from September 1918 from New York, the other from November 1918 from Alaska.
genetic variance
The result: while the Berlin genomes are very similar – as far as they can be judged by their incompleteness – the genomes of the virus isolated from the Munich sample actually show several differences. Genetic sequences from New York and Alaska show more mutations. Taken together, these comparisons show that there is measurable genetic variation, and that genomes from the same continents and during the same period of the epidemic showed lower overall divergence than genomes from different continents and during different time periods.
A comparison of genomes from the early and peak phase of the pandemic also provides information about mutations that could increase the virulence of the influenza strain at that time. “Our results show differences at two sites in the influenza A virus nucleoprotein gene,” the researchers report. “These are related to resistance to the host’s antiviral response.” While pre-pandemic virus strains still bear structures typical of avian influenza viruses at these sites, the 1918 and 1919 influenza viruses replaced these appendages that are still present in most people today. Pathogenic H1N1 strains. As a result of these mutations, the virus could have adapted better to humans.
Further analysis is required
In addition, the researchers performed molecular clock modeling to estimate the evolutionary time frames over which the virus evolved. Accordingly, the mutation rate of the virus may be higher than previously thought, making the pandemic virus the progenitor of all H1N1 influenza viruses currently in existence. “Our analyzes indicate that the seasonal H1N1 viruses circulating today evolved from the pandemic strain at that time,” the authors wrote. This contrasts with the hypotheses that today’s seasonal influenza viruses evolved primarily through the exchange of genome parts between different strains.
“Given the very small sample size–three complete genomes and two incomplete genomes–all results should be considered preliminary,” the authors say. In order to substantiate the hypotheses, further genome analyzes are required. “We anticipate that the main obstacle to a better understanding of the evolution of 1918 influenza viruses will be the identification of the remaining pathological samples,” they wrote. However, sick collections in museums can be an important resource.
Source: Livia Patrono (Robert Koch Institute, Berlin) et al., Nature Communications, doi: 10.1038/s41467-022-29614-9

“Alcohol buff. Troublemaker. Introvert. Student. Social media lover. Web ninja. Bacon fan. Reader.”





More Stories
“Time seems to cure long Covid.”
Science: The use of artificial intelligence is changing the way hospitals operate
Simple recipe: sweet cream cheese slices from the tray